GROMACS Tutorial
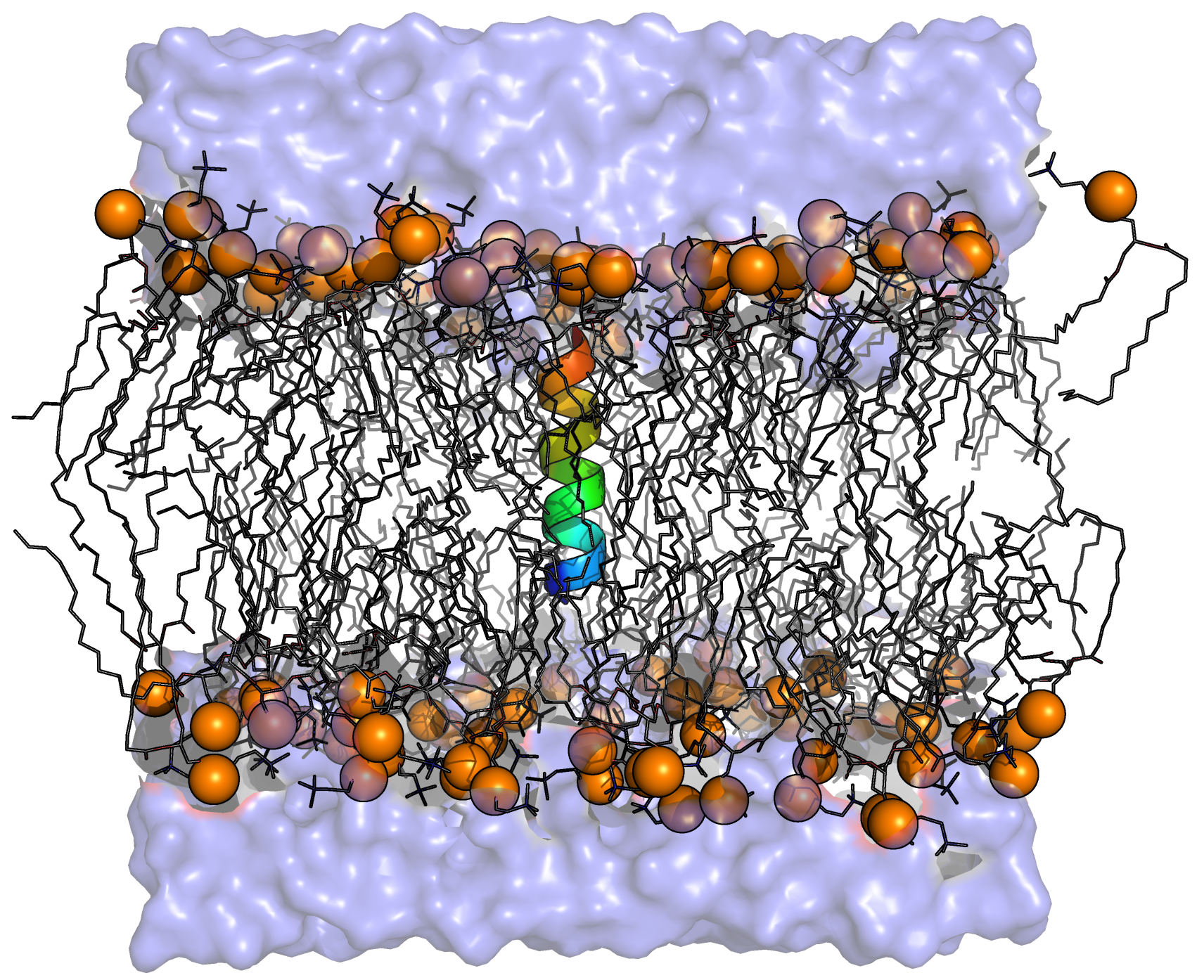
Membrane Protein: KALP15 in DPPC
Justin A. Lemkul, Ph.D.
Virginia Tech Department of Biochemistry
|
This tutorial will guide the user through the preparation and simulation of a simple membrane protein, in this case KALP15, in a model membrane, DPPC. The tutorial assumes the user has already successfully completed the Lysozyme tutorial, some other tutorial, or is otherwise well-versed in GROMACS simulation methods and topology organization. The level of detail in this tutorial will be focused on membrane protein-specific considerations, and will not provide an exhaustive explanation of every step, as with the Lysozyme tutorial. This tutorial assumes you are using GROMACS version 2018 or newer. Please note that the purpose of this tutorial is instructional, to build a membrane protein system and also to understand GROMACS force field organization and methods for modification. This tutorial is not an endorsement or suggestion that you use these specific parameters for your simulation. The approach taken here works well in the case where one needs to augment a parameter set with other parameters that have been derived in a consistent manner. Some force fields already include everything you need, without modification. For instance, it is unwise to try to literally follow this approach for a force field like CHARMM36, as it needs no modification. In that case, you are much better off building the system with CHARMM-GUI. As of summer 2023, the required Berger lipid parameters are no longer available online. The authors themselves have raised issues of their validity and they are not available. This tutorial will remain online as a reference until I rewrite it. Please DO NOT proceed and expect the tutorial to work. It will not. |

Site design and content copyright Justin Lemkul
Problems with the site? Send them to the Webmaster